23MALDI-TOF ANALYSIS
MALDI-TOF analysis
 The new Spectrum type experiment type provides a comprehensive platform for importing, preprocessing and summarizing spectrum fingerprints obtained by e.g. Matrix-Assisted Laser Desorption/Ionization Time-Of-Flight Mass Spectrometry (MALDI TOF MS), Liquid Chromatography – Mass Spectrometry (LC MS), ElectroSpray Ionization (ESI), etc.
The new Spectrum type experiment type provides a comprehensive platform for importing, preprocessing and summarizing spectrum fingerprints obtained by e.g. Matrix-Assisted Laser Desorption/Ionization Time-Of-Flight Mass Spectrometry (MALDI TOF MS), Liquid Chromatography – Mass Spectrometry (LC MS), ElectroSpray Ionization (ESI), etc.
The BioNumerics software offers an intuitive environment with easy-to-use workflow templates for importing, resampling and preprocessing spectral data. Starting from existing templates, the user can create customized workflows according to specific needs. Summarizing spectra and creating comparisons and identification projects is easy using BioNumerics' impressive set of analysis tools.
Processing spectrum data in BioNumerics
Importing spectrum fingerprints
The redesigned import window allows fast import of various spectrum data formats:
· Bruker's MALDI BioTyper® Main Spectra (MSP) from *.btmsp files.
· Spectrum data or peak tables from text files.
· mzML files (raw or processed spectrum data).
Preprocessing of spectra
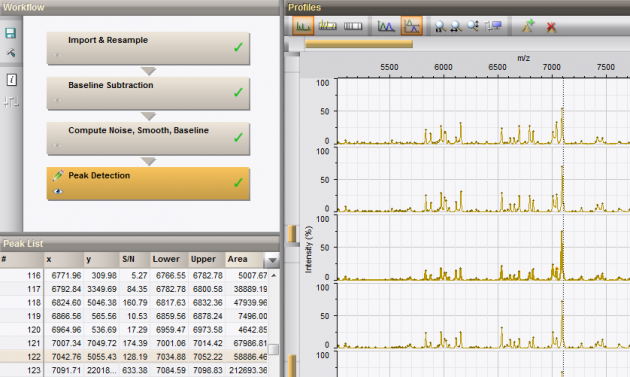
![]() provides fully automated workflow templates which enable easy preprocessing of raw spectrum fingerprints. The user can easily create customized templates starting from existing templates, with full control over the parameters in each action of the pipeline to suit specific needs.
provides fully automated workflow templates which enable easy preprocessing of raw spectrum fingerprints. The user can easily create customized templates starting from existing templates, with full control over the parameters in each action of the pipeline to suit specific needs.
· Fast importing & resampling actions.
· Comprehensive options for cleaning-up spectrum profiles using baseline subtraction, noise elimination and curve smoothing.
· Automatic peak calling with manual editing options.
Summarizing spectra
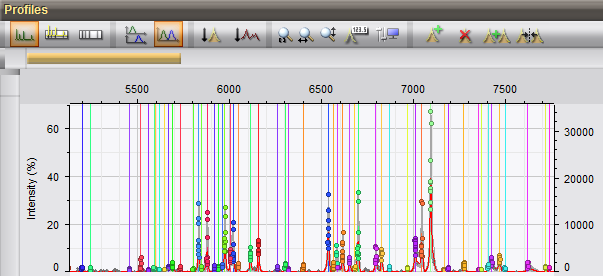
The creation of summary spectra is an essential step in MALDI-TOF analysis. The summarizing methods in BioNumerics are based on a peak matching and member average. Flexible settings can be applied to filter out spectrum fingerprints of low quality. Similarity values allow easy inspection of the coherence of included spectrum profiles.
Required modules:
Relevant applications:
